Bio-inorganic Chemistry
The nutrient trace metals are essential to life but are highly toxic in excess. A balance between deficiency and toxic excess must be maintained. The secrets of their catalytic and structural roles in enzymes are under intensive scrutiny.
Acquisition of the metals is mediated by molecular membrane pumps and by transport proteins which take the metal to its destinations in biological cells. Figure 1 shows a simplified representation of transport of copper in mammalian cells. Copper imbalance is associated with acquired and inherited diseases. These include major neurodegenerative diseases (Alzheimer, Parkinson, motor neurone, ‘mad cow’ (prion)) and copper deficiency and overload conditions (Menkes, Wilson). See, eg, ref [1]. Each of the key proteins associated with these diseases is a copper-binding protein. Very recently, copper transport proteins have been implicated as critical components of tuberculosis, of cancer cell proliferation and of fungal virulence.
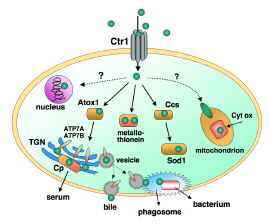
Figure 1. Over-simplified model of copper transport in human cells such as liver hepatocytes which receive much of the copper absorbed from the small intestine. Membrane pumps Ctr1 and ATP7A,B and chaperones Ccs and Atox1 transport CuI. Ceruloplasmin (Cp), Cytochrome c oxidase (Cyt ox) and superoxide dismutase (Sod1) are redox enzymes which employ the CuII/CuI couple. As well as loading Cp with copper, the trans-Golgi network (TGN) inserts copper into many enzymes. The ATP7B pump performs two functions: it transports nutrient copper into the TGN and excess copper out of the cell (by trafficking to the cell membrane via vesicles). It may also supply copper to phagosomes for destruction of invading bacteria. ATP7A is the equivalent pump in most other cells.
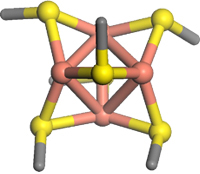
The molecular pump Ctr1 (Copper Transporter no. 1; Figure 1) is primarily responsible for import of copper into human cells. The yeast version appears to interact with the transport proteins (chaperones) via a beautiful Cu4S6 cluster (Figure 2).[2]
Figure 2. Cu4S6 cluster (Cu atoms are pink)
In addition, the cancer drug cis-platin enters certain cells via the Ctr1 pump. Interactions of cisplatin with the copper proteins appears to be a major cause of loss of drug and side effects. We are studying the pumps and chaperones and their interactions with cisplatin.[3]
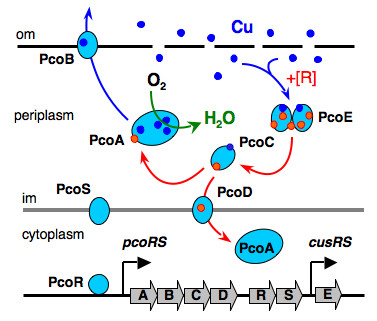
Certain bacteria have developed the unusual ability to survive in environments with millimolar concentrations of copper (>1,000 times higher than normal nutrient levels). They have evolved clusters of genes which are induced by copper and whose protein products combine to export excess copper (Figure 3).
Figure 3. Resistance proteins expressed to periplasm in E. Coli cells.
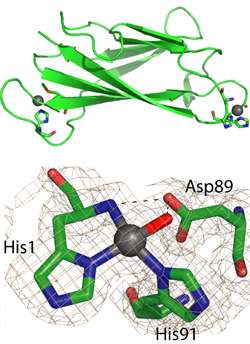
We have studied the properties of members of the cluster. In particular, the protein PcoE has the properties of a “metal sponge”, ie, it can bind multiple copper ions as an initial defence against elevated copper levels.[4] These copper ions may be then transferred to PcoC (CopC) which has the unique ability to bind copper in either of its oxidation states Cu(I) and Cu(II), one at either end of the molecule (Figure 4).[5]
Figure 4. Copper chaperone protein PcoC (CopC). (a) Cu atoms bind at each end; (b) Detail of CuIIbinding site.
In addition, the oxidase enzyme PcoA catalyses oxidation of Cu(I) bound to PcoC catalytically to the less toxic Cu(II) form (Figure 5).[6] This can then be pumped out of the cell. Interestingly, PcoA contains nutrient copper (Figure 3, 5) which has evolved to eliminate toxic excess copper!!
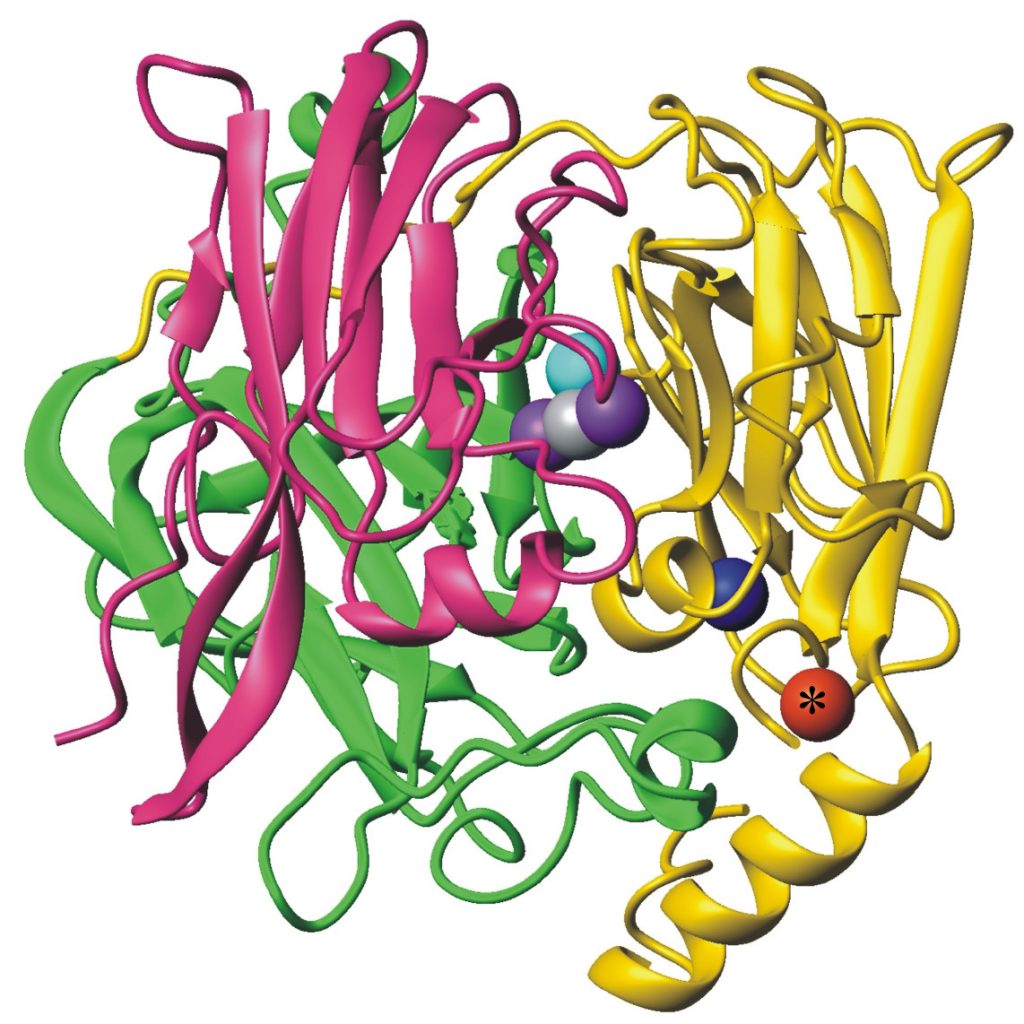
Figure 5. Cuprous oxidase enzyme CueO features multi-copper oxidase machinery (blue, teal, purple copper atoms) which reduces O2 to H2O and the oxidation site (red) which binds substrate Cu(I).
The protein CopK from C. metallidurans also binds two copper ions but with very different structural features (Figure 6; compare with PcoC: Figure 4). Remarkably, a unique cooperativity means that binding of CuI increases the affinity for CuII by six orders of magnitude. The CuII binding site is assembled upon insertion of CuI into its site.[7, 8]
Figure 6. Copper resistance protein CopK. Apo-protein (left) binds both Cu(I) (orange; internal site) and Cu(II) (blue; surface site) with major structural rearrangement to give the holo-protein (right).
We have commenced work on the membrane pump CopA (related to the Menkes disease protein: ATP7A of Figure 1) and its biological partner CueO.[6] The inner membrane pump PcoD from E. coli (see Figure 3) is also under study.
The proteins are generated via molecular genetics and then purified. The molecular probes needed are provided by techniques such as NMR, ESR, MS, fluorescence, X-ray crystallography, electro-chemistry and quantitative HPLC.
New projects include:
- design and synthesis of new chromophoric ligands to act as quantitative probes of bio-metals, both in vivo and in vitro;[9, 10]
- Cell-free expression of membrane proteins (such as CopA (ATP7A of Figure 1) and PcoD (see Figure 3).
Selected Publications:
- M W. Brazier, A. G. Wedd et al, “Metal attenuating therapies in neurodegenerative disease“, Expert Reviews of Neurotherapeutics 2011, 11, 1-29.
- Z. Xiao, F. Loughlin, G. N. George, G. Howlett and A.G. Wedd. “The C-terminal domain of the membrane copper transporter Ctr1 from Saccharomyces cerevisiae binds four Cu(I) ions as a cuprous-thiolate polynuclear cluster. Sub-femtomolar Cu(I) affinity of three proteins involved in copper trafficking”, J. Amer. Chem. Soc. 2004, 126, 3801-3890.
- C. M. Sze, G. N. Khairallah, Z. Xiao, P. S. Donnelly, R. A. J. O’Hair and A.G. Wedd. “Interaction of Cisplatin and Analogues with a Met-Rich Protein Site”, J. Biol. Inorg. Chem. 2009, 14, 163-165.
- M. Zimmermann, S. Udegadara, C. M. Sze, T. Ryan, Z. Xiao and A. G. Wedd, “PcoE, a metal sponge in the periplasm of copper resistant E. coli”, J. Inorg. Biochem. 2012, 17, in press (Hans Freeman Memorial Issue).
- L. Zhang, M. Koay, M. J. Maher, Z. Xiao and A. G. Wedd. “Intermolecular Transfer of Copper Ions from the CopC Protein of Pseudomonas syringae. Crystal Structures of Fully-Loaded CuICuII Forms.”, J. Am. Chem. Soc. 2006, 128, 5834-5850.
- K. Y. Djoko, L. X. Chong, Z. Xiao and A. G. Wedd, “Reaction Mechanisms of the Multicopper Oxidase CueO from Escherichia coli Support its Functional Role as a Cuprous Oxidase”, J. Amer. Chem. Soc. 2010, 132, 2005-2015.
- L. X. Chong, M. R. Ash, M. J. Maher, M. G. Hinds, Z. Xiao and A. G. Wedd. “Unprecedented Binding Cooperativity between Cu(I) and Cu(II) in the Copper Resistance Protein CopK from Cupriavidus metallidurans CH34. Implications from Structural Studies by NMR Spectroscopy and X-ray Crystallography.”, J. Am. Chem. Soc. 2009, 13, 3564-3579.
- M.-R. Ash, L. X. Chong, M. J. Maher, M. G. Hinds, Z. Xiao and A. G. Wedd, “Molecular Basis of the Cooperative Binding of Cu(I) and Cu(II) to the CopK Protein from Cupriavidus metallidurans CH34”, Biochem. 2011, 50, 9237-9247.
- Z. Xiao, J. Brose, S. Schimo, S. M. Ackland, S. La Fontaine and A. G. Wedd, “Unification of the copper(I) binding affinities of the human copper metallo-chaperone Atox1 and related proteins: detection probes and affinity standards”, J. Biol. Chem. 2011, 286, 11047-11055.
- Z K. A. Price, J. L. Hickey, Z. Xiao, A. G. Wedd, S. A. James, J. R. Liddell, P. J. Crouch, A. R. White and P. S. Donnelly, “The Challenges of Using Fluorescent Sensor CS1 to Track Intracellular Cu”, Chemical Science, 2012, in proof (SC-EDG-03-2012-020397.R1).